Blast compare two protein sequences9/28/2023  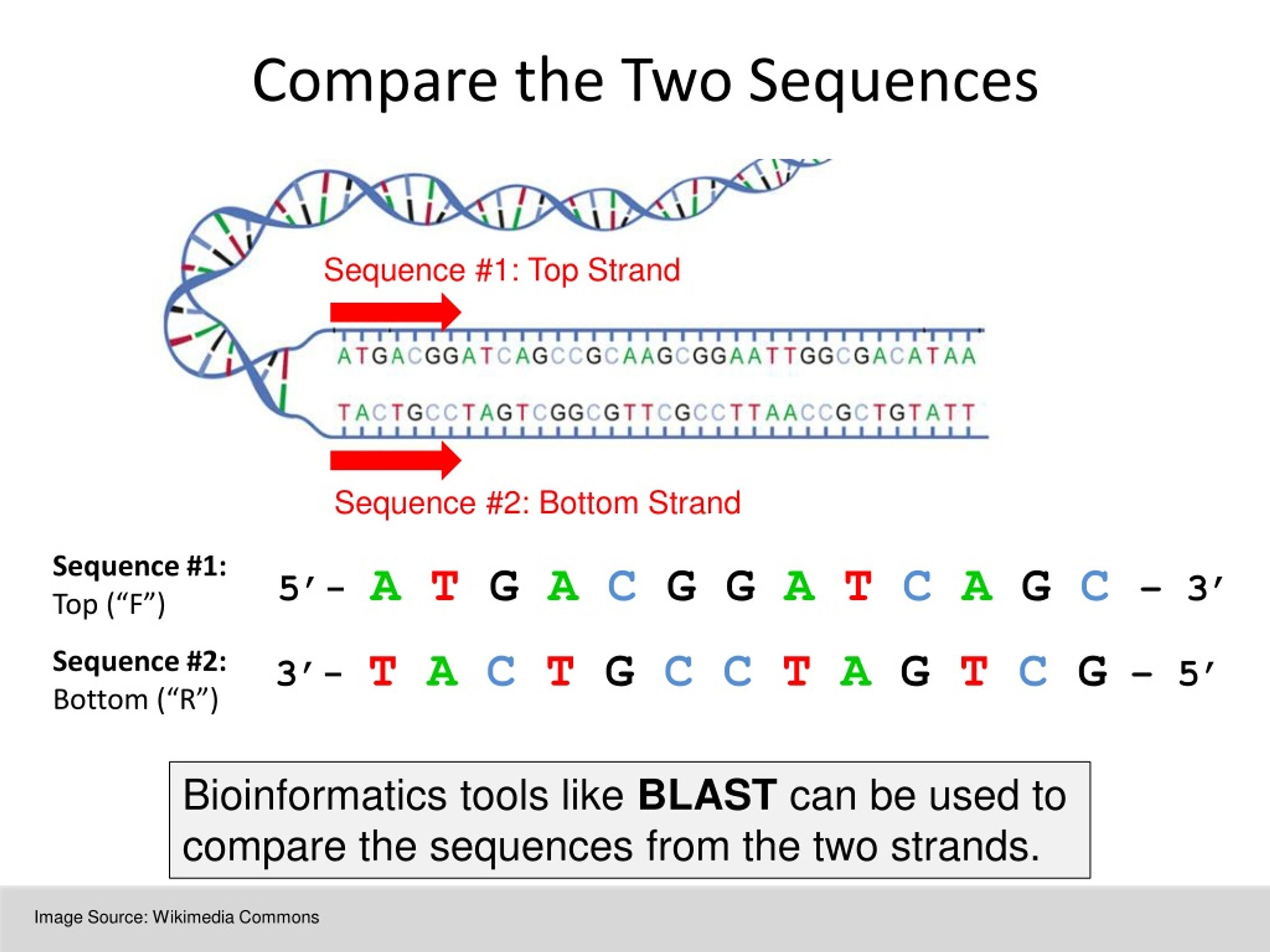
The sequence is broken down into words of a fixed length (usually three aminoacids or eleven DNA bases).Your input sequence is analyzed and low-complexity regions (think of repeating patterns) that do not add meaningful information to the search are stripped.Compared with those older approaches, BLAST takes a couple of steps to significantly reduce the complexity of the problem: If you scale this problem to thousands of sequences you will soon find that, regardless of the computational power you put into it, it will quickly become unfeasible. I encourage you to check out the reference for yourself, but in the meantime let’s take a quick look at how it works and what makes it so fast.īefore BLAST, an exhaustive comparison between two sequences would take a relatively long time to perform. The BLAST algorithms were first published by Altschul et al. While it doesn’t promise always to yield an optimal result, it can be orders of magnitude faster than the previous alternatives. Unlike the algorithms that came before, BLAST uses a heuristic approach. What is BLAST and why is it useful?īLAST is a family of algorithms designed for retrieving sets of data, similar to query strings, from a significantly large body of data. That’s where tools like BLAST come into play. This proves particularly useful when researches get ahold of newly sequenced data, as it is important to be able to check if it looks similar to other already known sequences.īecause the current corpus of known data is extremely big, we need specialised tools to search it efficiently and to measure the degree of similarity between sets of results. Today, thousands of genomes from different species have been sequenced and are available for us to work with, revealing valuable information about their genes and their corresponding function. That usually holds true and this is the basis for most evolutionary and phylogenetic analyses. One might expect that genes between two species that are closely related would look more similar than genes from species that had deviated from each other a long time ago. When species start to diverge from each other, due to migration, a natural event or an evolutionary constraint, their gene pools tend to accumulate different mutations over time. Within biology, two genes are defined as homologues if they come from a common source of ancestry i.e. a medium (4GB RAM or bigger) Exoscale instance running Ubuntu 16.04 LTS.to be comfortable working in a Linux terminal.to know how to create a new Exoscale instance and how to to connect to it.How to perform a query against a precomputed database of sequences.How to install BLAST on a fresh Ubuntu 16.04 LTS instance.The theory behind BLAST, one of the most widely-deployed bioinformatic algorithms.In this series of posts, I’m going to introduce you to some of the bioinformatics tools and techniques that field biologists, such as myself, use in our daily work. Bioinformatics is a huge part of modern biological study. 
0 Comments
Leave a Reply.AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
